Ribosomes: Cast of characters
Ribosomes play a central role in protein synthesis and catalyses formation of the peptide bonds that link amino acids into a polypeptide.
Only polypeptides with ER signal sequences can be inserted into or across the ER membrane as their amalgamation proceeds.
The idea that the ER is involved in transporting newly synthesised proteins to various locations within the cell first emerged from studies by Colvin Redman and David Sabatini of protein synthesis in isolated vesicles of rough ER (ER vesicles with attached ribosomes).
After briefly incubating rough ER vesicles in the presence of radioactive amino acids and the other components required for protein synthesis, they added the antibiotic puromycin to halt the process. In addition to blocking further protein synthesis, puromycin causes the partially completed polypeptide chains to be released from the ribosomes.
When the ribosomes and membrane vesicles were then separated and analysed to see where the newly made, radioactive polypeptide chains were located, a substantial fraction of the radioactivity was found inside the ER lumen. Such results suggested that some newly forming polypeptides begin to pass into the lumen of the ER as they are being synthesised, allowing them to be subsequently routed through the lumen of the ER to their correct destinations.
Advertisement
If some polypeptides move directly into the lumen of the ER as they are being synthesised, how does the cell determine which polypeptides are to be handled in this way? An answer was first suggested in 1971 by Giinter Blobel and David Sabatini, whose model was called the signal hypothesis because it proposed that some sort of intrinsic molecular signal distinguishes such polypeptides from the many polypeptides destined to be released into the cytosol.
This hypothesis has had such a profound impact on evidence that proteins synthesised on ribosomes attached to ER membranes pass directly into the ER lumen. ER vesicles containing attached ribosomes were isolated and incubated with radioactive amino acids to label newly made, polypeptide chains.
Next, protein synthesis was halted by adding puromycin, which also causes the newly forming polypeptide chains to be released from the ribosomes. The ribosomes were then removed from the membrane vesicles and the amount of radioactive protein associated with the ribosomes and in the membrane vesicles was measured.
The graph shows that after the addition of puromycin, radioactivity is lost from the ribosomes and appears inside the vesicles, suggesting that the newly forming polypeptide chains are inserted through the ER membrane as they are being synthesised, and puromycin causes the chains to be prematurely released into the vesicle lumen.
The field of cell biology that Blobel was awarded the Nobel Prize in 1999 for his many years of work in demonstrating the validity and widespread relevance of the idea that proteins have intrinsic signals that govern their transport and localisation within the cell. The signal hypothesis stated that for polypeptides destined for the ER, the first segment of the polypeptide to be synthesised, the N-terminus, contains an ER signal sequence that directs the ribosome-mRNA-polypeptide complex to the surface of the rough ER, where the complex anchors at a protein “dock” on the ER surface.
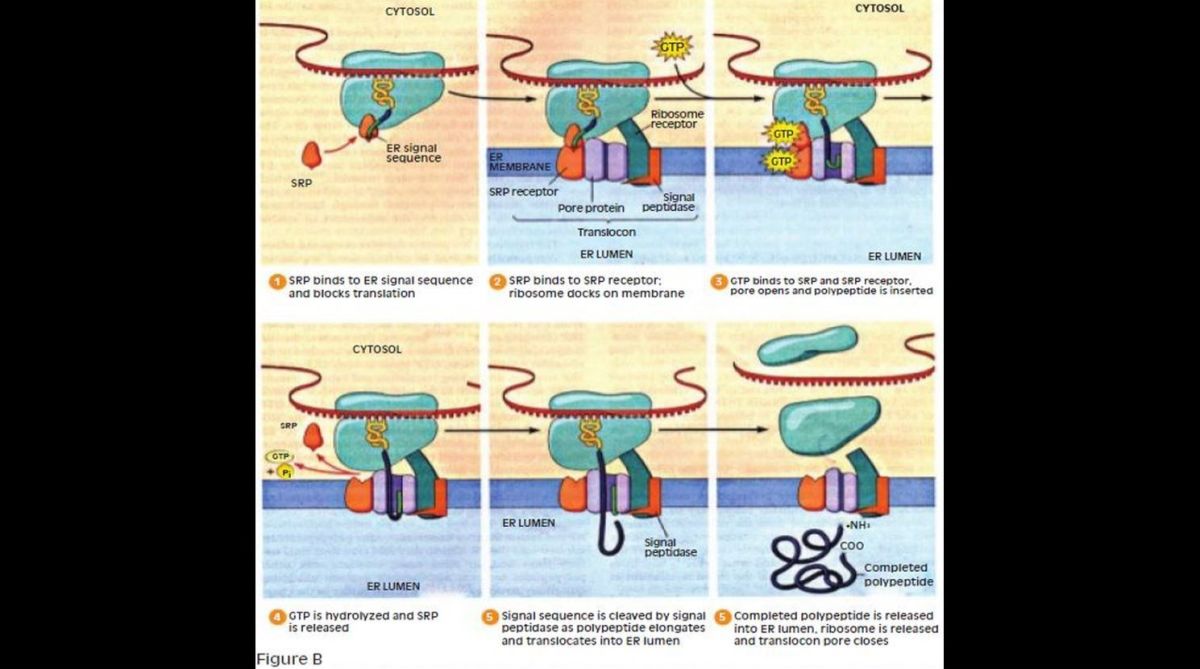
Then, as the polypeptide chain elongates during mRNA translation, it progressively crosses the ER membrane and enters the ER lumen. Figure B shows a current model for the signal mechanism. It is now well established that the growing polypeptide translocates through a hydrophilic pore created by one or more membrane proteins. The complex of membrane proteins that carry out translocation is called the translocon.
Evidence for the actual existence of ER signal” sequences was obtained shortly thereafter by Cesar Milstein and his associates, who were studying the synthesis of the small subunit, or light chain, of the protein immunoglobulin G.
In cell-free systems containing purified ribosomes and the components required for protein synthesis, the mRNA coding for the immunoglobulin light chain directs the synthesis of a polypeptide product that is 20 amino acids longer at its N-terminal end than the authentic light chain itself.
Adding ER membranes to this system leads to the production of an immunoglobulin light chain of the correct size. Such findings suggest that the extra 20-amino acid segment is functioning as an ER signal sequence, and that this signal sequence is removed when the polypeptide moves into the ER.
Subsequent studies revealed that other polypeptides destined for the ER also possess an N-terminal sequence that is required for targeting the protein to the ER and that is removed as the polypeptide moves into the ER. Proteins containing such ER signal sequences at their N-terminus are often referred to as preproteins (e.g., prelysozyme, preproinsulin, pretrypsinogen, and so forth).
Sequencing studies have revealed that the amino acid compositions of ER signal sequences are surprisingly variable, but several unifying features have been noted. ER signal sequences are typically 15 to 30 amino acids long and consist of three domains: a positively charged N-terminal region, a central hydrophobic region, and a polar region adjoining the site where cleavage from the mature protein will take place.
The positively charged end may promote interaction with the hydrophilic exterior of the ER membrane, and the hydrophobic region may facilitate interaction of the signal sequence with the membrane’s lipid interior. In any case, it is now established that only polypeptides with ER signal sequences can be inserted into or across the ER membrane as their synthesis proceeds.
In fact, when recombinant DNA methods are used to add ER signal sequences to polypeptides that do not usually have them, the recombinant polypeptides are directed to the ER.
The writer is associate professor and head, department of botany, Ananda Mohan College.
Advertisement